NeuArc2Fun project: "Biological neural networks: from structure to function"
Summary:
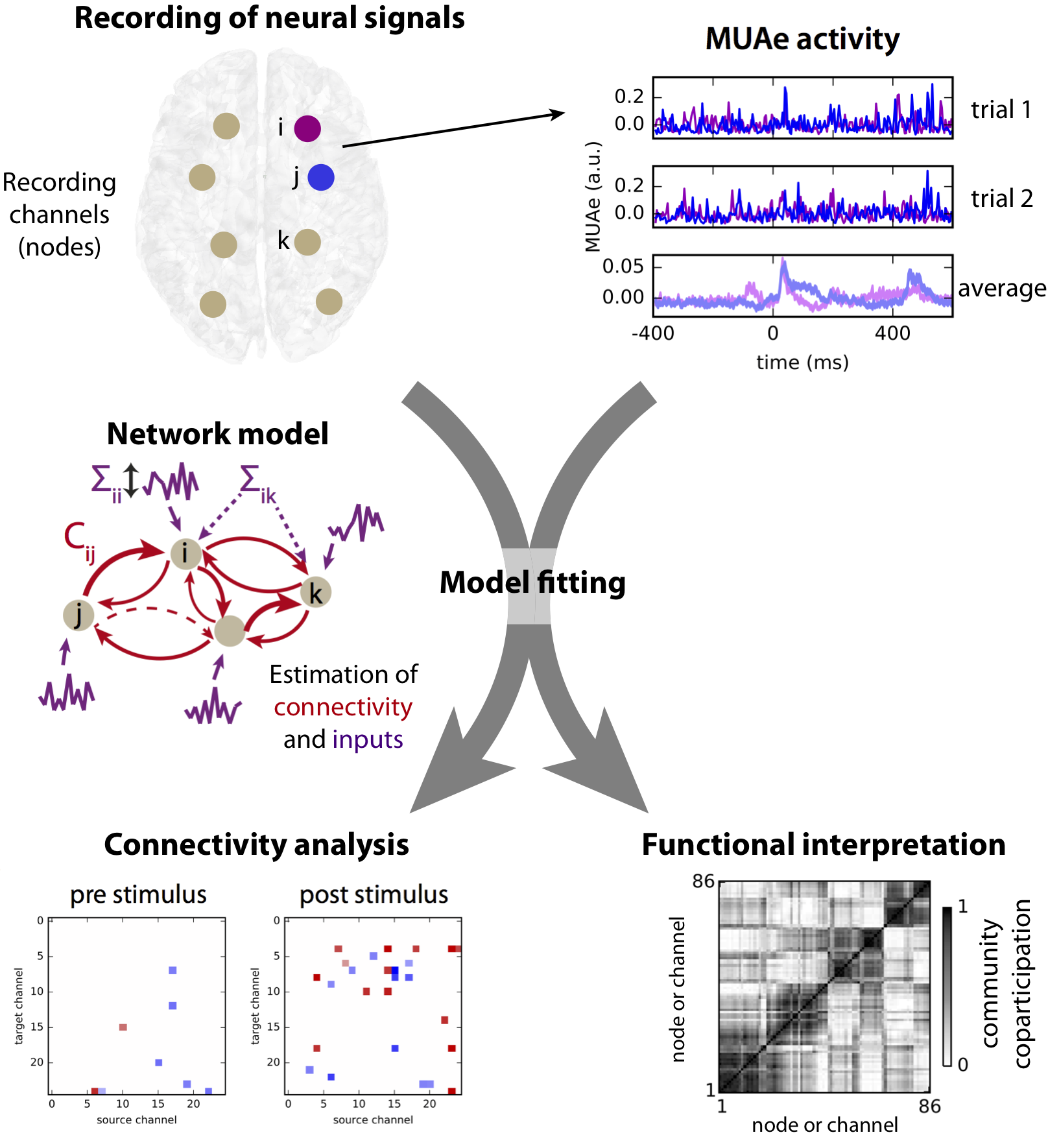
The NeuArc2Fun project proposal was to set at the connection between theoretical and experimental neuroscience. The goal was the investigation of information processing in neural networks with feedback. Technically, it aimed to adapt and improve analysis tools for newly collected data by nowadays recording and imaging techniques, which have much improved recently. The whole approach relies on the estimation and interpretation of interactions between large (mesoscopic) populations of neurons from their activity that is simultaneously recorded. This includes electrophysiology (e.g., Utah electrode array), calcium imaging or fMRI (considering whole-brain activity). In particular, the focus of NeuArc2Fun was on “large” networks (i.e., from 20 to 100 of nodes), which is suitable for state-of-the-art measurements. This requires novel analysis tools of recurrently connected neural networks, in order to bridge the structural and functional levels (from connectivity to patterns of correlated activity and information processing).
The key for a useful formalism is finding the adequate balance between the mathematical tractability and biological realism of the model. To address this trade-off problem, NeuArc2Fun focuses on the mesoscopic level, i.e., scales at which many interacting neural populations can be simultaneously recorded by current state-of-the-art experimental techniques, such as electrode arrays. The advantage of this model-based approach is the ability to make predictions about the role of each component of the model –in particular, its heterogeneous connectivity– in shaping neural activity. A particular focus was on bridging several disciplines in a common comprehensive formalism: dynamic system, graph theory, statistics and information theory.
The primary application of the framework was targeted at electrophysiological data recorded in monkeys from the laboratory of Prof. Thiele in Newcastle University. Modeling such data is particularly challenging because they exhibit a large variability over repeated trials in the same condition, which hinders the extraction of consistent condition-specific information.
Main publications and results:
-
Gilson M*, Tauste Campo A*, Chen X, Thiele A, Deco G (in press)
Non-parametric test for connectivity detection in multivariate autoregressive networks and application to multiunit activity data.
Network Neurosci
doi: 10.1162/NETN_a_00019
-
Python code to compare the method with Granger causality: MVAR_Granger_detection.py
-
Gilson M, Kouvaris NE, Deco G, Zamora-López G (accepted)
Novel framework to analyze complex network dynamics.
Phys Rev E
arxiv preprint
-
Gilson M (in press)
Analysis of fMRI data using noise-diffusion network models: a new covariance-coding perspective.
Biol Cybern
online first; biorxiv preprint

